Detection Limits
This guide shows how to calculate detection limits, in mass, or in photometry, as a function of orbital parameters for different combinations of data.
There are a few use cases for this:
- Mass limit vs semi-major axis given one or more images and/or contrast curves
- Mass limit vs semi-major axis given an RV non-detection
- Mass limit vs semi-major axis given proper motion anomaly from the GAIA-Hipparcos Catalog of Accelerations
- Any combination of the above
We will once more use some sample data from the system HD 91312 A & B discovered by SCExAO.
using Octofitter
using OctofitterImages
using OctofitterRadialVelocity
using Distributions
using Pigeons
using CairoMakie
using PairPlotsPhotometry Model
We will need to decide on an atmosphere model to map image intensities into mass. Here we use the Sonora Bobcat cooling and atmosphere models which will be auto-downloaded by Octofitter:
const cooling_tracks = Octofitter.sonora_cooling_interpolator()
const sonora_temp_mass_L = Octofitter.sonora_photometry_interpolator(:Keck_L′)(::Octofitter.var"#model_interpolator#450"{Interpolations.FilledExtrapolation{Float64, 2, Interpolations.ScaledInterpolation{Float64, 2, Interpolations.BSplineInterpolation{Float64, 2, Matrix{Float64}, Interpolations.BSpline{Interpolations.Linear{Interpolations.Throw{Interpolations.OnGrid}}}, Tuple{Base.OneTo{Int64}, Base.OneTo{Int64}}}, Interpolations.BSpline{Interpolations.Linear{Interpolations.Throw{Interpolations.OnGrid}}}, Tuple{StepRangeLen{Float64, Base.TwicePrecision{Float64}, Base.TwicePrecision{Float64}, Int64}, StepRangeLen{Float64, Base.TwicePrecision{Float64}, Base.TwicePrecision{Float64}, Int64}}}, Interpolations.BSpline{Interpolations.Linear{Interpolations.Throw{Interpolations.OnGrid}}}, Float64}, Float64, Float64, Float64, Float64}) (generic function with 1 method)Proper Motion Anomaly Data
We start by defining and sampling from a model that only includes proper motion anomaly data from the HGCA:
B = Planet(
name="B",
basis=Visual{KepOrbit},
observations=[],
variables=@variables begin
a ~ LogUniform(1, 65)
e ~ Uniform(0,0.9)
ω ~ Uniform(0,2pi)
i ~ Sine() # The Sine() distribution is defined by Octofitter
Ω ~ Uniform(0,pi)
mass = system.M_sec
θ ~ Uniform(0,2pi)
tp = θ_at_epoch_to_tperi(θ, 57423.0; M=system.M, e, a, i, ω, Ω) # epoch of GAIA measurement
end
)
HD91312_pma = System(
name="HD91312_pma",
companions=[B],
observations=[HGCAInstantaneousObs(gaia_id=6166183842771027328)],
variables=@variables begin
M_pri ~ truncated(Normal(0.95, 0.05), lower=0.1) # Msol
M_sec ~ LogUniform(0.2, 65) # MJup
M = M_pri + M_sec*Octofitter.mjup2msol # Msol
plx ~ gaia_plx(gaia_id=6166183842771027328)
# Priors on the center of mass proper motion
pmra ~ Normal(0, 1000)
pmdec ~ Normal(0, 1000)
end
)
model_pma = Octofitter.LogDensityModel(HD91312_pma)LogDensityModel for System HD91312_pma of dimension 11 and 4 epochs with fields .ℓπcallback and .∇ℓπcallback
Sample:
using Pigeons
chain_pma, pt = octofit_pigeons(model_pma, n_chains=16, n_chains_variational=16, n_rounds=12);[ Info: Sampler running with multiple threads : true
[ Info: Likelihood evaluated with multiple threads: false
┌ Info: Starting values not provided for all parameters! Guessing starting point using global optimization:
│ num_params = 11
└ num_fixed = 0
┌ Info: Found sample of initial positions
│ logpost_range = (-27.525410151437285, -15.472239835075007)
└ mean_logpost = -17.23899154539307
────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
scans restarts Λ Λ_var time(s) allc(B) log(Z₁/Z₀) min(α) mean(α) min(αₑ) mean(αₑ)
────────── ────────── ────────── ────────── ────────── ────────── ────────── ────────── ────────── ────────── ──────────
2 0 3.94 2.15 0.234 4.15e+06 -1.66e+08 0 0.804 1 1
4 0 4.52 3.5 0.0728 1.45e+05 -8.06e+06 0 0.741 1 1
8 0 4.17 4.62 0.143 2.68e+05 -4.43e+05 0 0.717 1 1
16 0 4.51 5.62 0.283 4.37e+05 -2.73e+05 0 0.673 1 1
32 0 5.32 5.9 0.559 7.8e+05 -9.5e+03 0 0.638 1 1
64 1 6.02 2.99 1.29 4.33e+07 -15.8 0 0.709 1 1
128 7 6.74 3.28 2.14 7.33e+07 -17 0 0.677 1 1
256 20 6.99 2.98 4.35 1.46e+08 -17.1 0 0.678 1 1
512 44 7.53 3.54 8.74 2.93e+08 -17.3 2.73e-37 0.643 1 1
1.02e+03 95 7.88 3.08 17.4 5.81e+08 -17.3 0.00179 0.647 1 1
2.05e+03 201 8.43 2.95 35.1 1.16e+09 -17.2 0.000102 0.633 1 1
4.1e+03 327 8.77 3.62 70.6 2.33e+09 -17.1 0.00241 0.6 1 1
────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────Plot the marginal mass vs. semi-major axis posterior with contours using PairPlots.jl:
pairplot(
PairPlots.Series(
(;
sma=log.(chain_pma[:B_a][:],),
mass=log.(chain_pma[:B_mass][:]),
),
label="PMA",
color=Makie.wong_colors()[1],
)=>(
PairPlots.Scatter(markersize=3,alpha=0.35),
PairPlots.Contour(sigmas=[1,3]),
PairPlots.MarginStepHist(),
),
labels=Dict(
:sma=>"log Semi-major axis [au]",
:mass=>"log Mass [Mⱼᵤₚ]"
)
)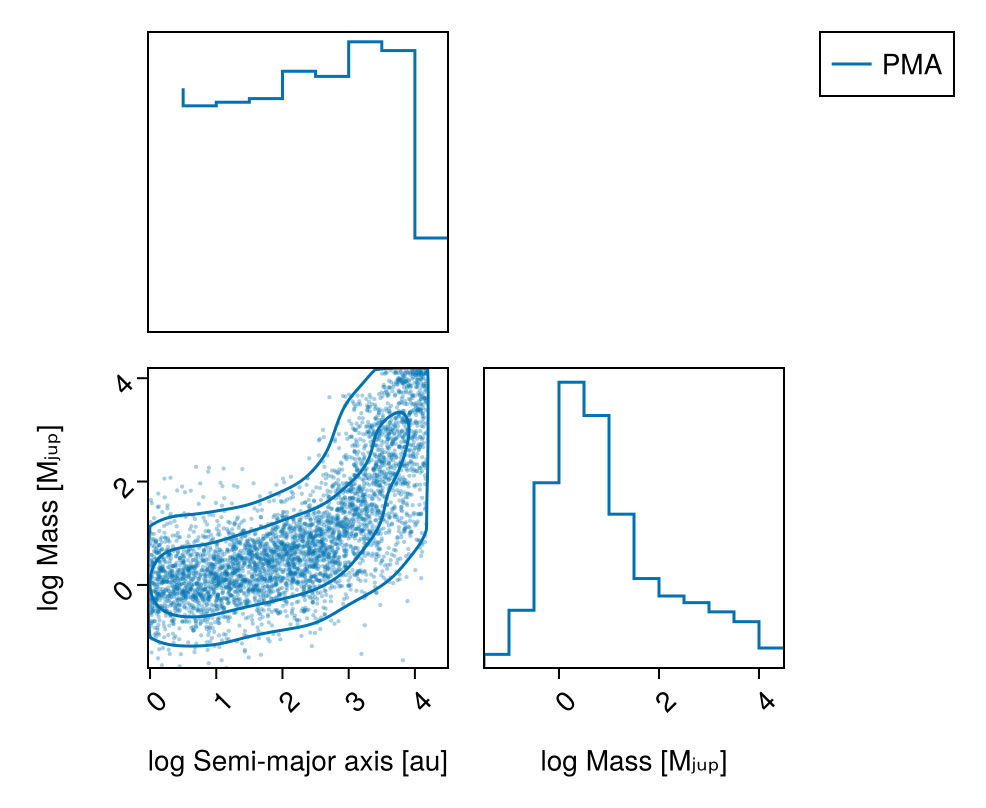
Image Data
using AstroImages
# download(
# "https://github.com/sefffal/Octofitter.jl/raw/main/docs/image-examples-1.fits",
# "image-examples-1.fits"
# )
# Or multi-extension FITS (this example)
image = AstroImages.load("image-examples-1.fits").*2e-7 # units of contrast
img_dat_table = Table([
(image=AstroImages.recenter(image), platescale=4.0, epoch=57423.6),
])
image_data = ImageObs(
img_dat_table,
name="imgdat-sim",
variables=@variables begin
# Planet flux in image units -- could be contrast, mags, Jy, or arb. as long as it's consistent with the units of the data you provide
flux = planet.L
# The following are optional parameters for marginalizing over instrument systematics:
# Platescale uncertainty multiplier [could use: platescale ~ truncated(Normal(1, 0.01), lower=0)]
platescale = 1.0
# North angle offset in radians [could use: northangle ~ Normal(0, deg2rad(1))]
northangle = 0.0
end
)OctofitterImages.ImageObs Table with 5 columns and 1 row:
image platescale epoch contrast ⋯
┌───────────────────────────────────────────────────────────────────
1 │ [-6.80461e-8 -6.247… 4.0 57423.6 65-element extrapol… ⋯B = Planet(
name="B",
basis=Visual{KepOrbit},
observations=[image_data],
variables=@variables begin
a ~ LogUniform(1, 65)
e ~ Uniform(0,0.9)
ω ~ Uniform(0,2pi)
i ~ Sine() # The Sine() distribution is defined by Octofitter
Ω ~ Uniform(0,pi)
mass = system.M_sec
# Calculate planet temperature from cooling track and planet mass variable
tempK = $cooling_tracks(system.age, mass)
# Calculate absolute magnitude
abs_mag_L = $sonora_temp_mass_L(tempK, mass)
# Deal with out-of-grid values by clamping to grid max and min
abs_mal_L′ = if isfinite(abs_mag_L)
abs_mag_L
elseif mass > 10
8.2 # jump to absurdly bright
else
16.7 # jump to absurdly dim
end
# Calculate relative magnitude
rel_mag_L = abs_mal_L′ - system.rel_mag + 5log10(1000/system.plx)
# Convert to contrast (same units as image)
L = 10.0^(rel_mag_L/-2.5)
θ ~ Uniform(0,2pi)
tp = θ_at_epoch_to_tperi(θ, 57423.6; M=system.M, e, a, i, ω, Ω)
end
)
HD91312_img = System(
name="HD91312_img",
companions=[B],
observations=[],
variables=@variables begin
# age ~ truncated(Normal(40, 15),lower=0, upper=200)
age = 10
M_pri ~ truncated(Normal(0.95, 0.05), lower=0.1) # Msol
# Mass of secondary
# Make sure to pick only a mass range that is covered by your models
M_sec ~ LogUniform(0.55, 65) # MJup
M = M_pri + M_sec*Octofitter.mjup2msol # Msol
plx ~ gaia_plx(gaia_id=6166183842771027328)
# Priors on the center of mass proper motion
# pmra ~ Normal(0, 1000)
# pmdec ~ Normal(0, 1000)
rel_mag = 5.65
end
)
model_img = Octofitter.LogDensityModel(HD91312_img)LogDensityModel for System HD91312_img of dimension 9 and 1 epochs with fields .ℓπcallback and .∇ℓπcallback
using Pigeons
chain_img, pt = octofit_pigeons(model_img, n_chains=5, n_chains_variational=5, n_rounds=7)(chain = MCMC chain (128×26×1 Array{Float64, 3}), pt = PT(checkpoint = false, ...))Plot mass vs. semi-major axis posterior:
vis_layers = (
PairPlots.Contour(sigmas=[1,3]),
PairPlots.MarginStepHist(),
)
pairplot(
PairPlots.Series(
(;
sma=log.(chain_pma[:B_a][:],),
mass=log.(chain_pma[:B_mass][:]),
),
label="PMA",
color=Makie.wong_colors()[1],
)=>vis_layers,
PairPlots.Series(
(;
sma=log.(chain_img[:B_a][:],),
mass=log.(chain_img[:B_mass][:]),
),
label="IMG",
color=Makie.wong_colors()[2],
)=>vis_layers,
labels=Dict(
:sma=>"log Semi-major axis [au]",
:mass=>"log Mass [Mⱼᵤₚ]"
)
)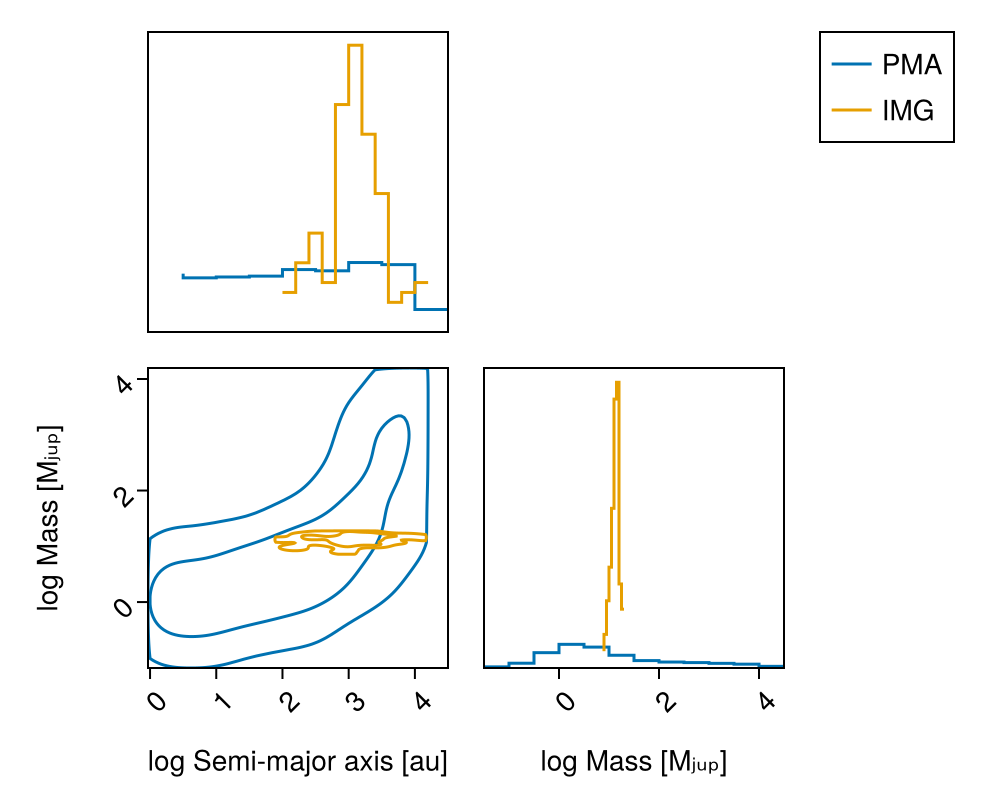
Image and PMA data
B = Planet(
name="B",
basis=Visual{KepOrbit},
observations=[image_data],
variables=@variables begin
a ~ LogUniform(1, 65)
e ~ Uniform(0,0.9)
ω ~ Uniform(0,2pi)
i ~ Sine() # The Sine() distribution is defined by Octofitter
Ω ~ Uniform(0,pi)
mass = system.M_sec
# Calculate planet temperature from cooling track and planet mass variable
tempK = $cooling_tracks(system.age, mass)
# Calculate absolute magnitude
abs_mag_L = $sonora_temp_mass_L(tempK, mass)
# Deal with out-of-grid values by clamping to grid max and min
abs_mal_L′ = if isfinite(abs_mag_L)
abs_mag_L
elseif mass > 10
8.2 # jump to absurdly bright
else
16.7 # jump to absurdly dim
end
# Calculate relative magnitude
rel_mag_L = abs_mal_L′ - system.rel_mag + 5log10(1000/system.plx)
# Convert to contrast (same units as image)
L = 10.0^(rel_mag_L/-2.5)
# L ~ Uniform(0,1)
θ ~ Uniform(0,2pi)
tp = θ_at_epoch_to_tperi(θ, 57423.6; M=system.M, e, a, i, ω, Ω)
end
)
HD91312_both = System(
name="HD91312_both",
companions=[B],
observations=[HGCAInstantaneousObs(gaia_id=6166183842771027328)],
variables=@variables begin
# age ~ truncated(Normal(40, 15),lower=0, upper=200)
age = 10
M_pri ~ truncated(Normal(0.95, 0.05), lower=0.1) # Msol
# Mass of secondary
# Make sure to pick only a mass range that is covered by your models
M_sec ~ LogUniform(0.55, 65) # MJup
M = M_pri + M_sec*Octofitter.mjup2msol # Msol
plx ~ gaia_plx(gaia_id=6166183842771027328)
# Priors on the center of mass proper motion
pmra ~ Normal(0, 1000)
pmdec ~ Normal(0, 1000)
rel_mag = 5.65
end
)
model_both = Octofitter.LogDensityModel(HD91312_both)
using Pigeons
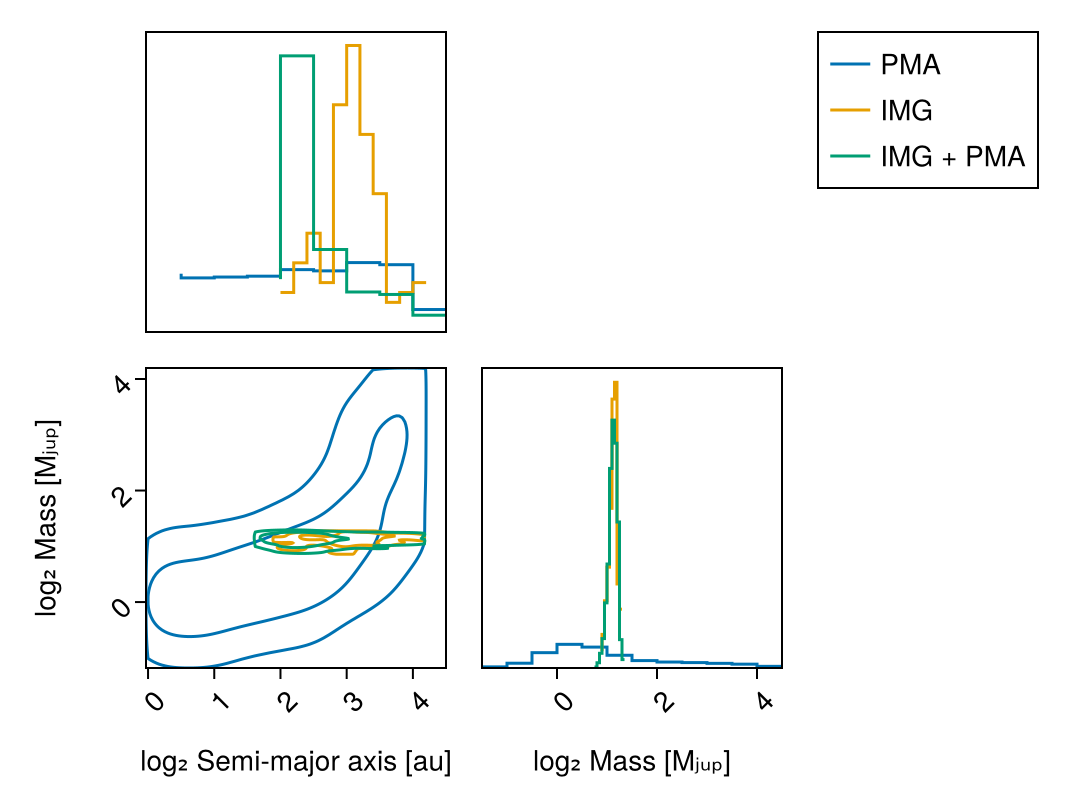
chain_both, pt = octofit_pigeons(model_both,n_chains=5,n_chains_variational=5,n_rounds=10)(chain = MCMC chain (1024×28×1 Array{Float64, 3}), pt = PT(checkpoint = false, ...))Compare all three posteriors to see limits:
vis_layers = (
PairPlots.Contour(sigmas=[1,3]),
PairPlots.MarginStepHist(),
)
pairplot(
PairPlots.Series(
(;
sma=log.(chain_pma[:B_a][:],),
mass=log.(chain_pma[:B_mass][:]),
),
label="PMA",
color=Makie.wong_colors()[1],
)=>vis_layers,
PairPlots.Series(
(;
sma=log.(chain_img[:B_a][:],),
mass=log.(chain_img[:B_mass][:]),
),
label="IMG",
color=Makie.wong_colors()[2],
)=>vis_layers,
PairPlots.Series(
(;
sma=log.(chain_both[:B_a][:],),
mass=log.(chain_both[:B_mass][:]),
),
label="IMG + PMA",
color=Makie.wong_colors()[3],
)=>vis_layers,
labels=Dict(
:sma=>"log₂ Semi-major axis [au]",
:mass=>"log₂ Mass [Mⱼᵤₚ]"
)
)